Setup Hayes PROCESS in R (4 Steps)
The following shows steps of setup Hayes Mediation PROCESS in R.
- Click here to visit the offcial Hayes PROCESS R package webpage. Then, click “Download PROCESS v 4.3.”

- Open the downloaded zip file.
You should be able to find the folder of “PROCESS v4.3 for R.” Within the folder, you should be able to find the “process.R ” file.
” file. - Open “process.R”
Assuming that you can R Studio in your computer, you can double-click the process.R . R Studio will open the process.R as an independent R file tap.
. R Studio will open the process.R as an independent R file tap. - Run “process.R”
The process.R code is very long. In order to select all its R code, right click the computer mouse and then click “Select all.” Then, hit Run in RStudio.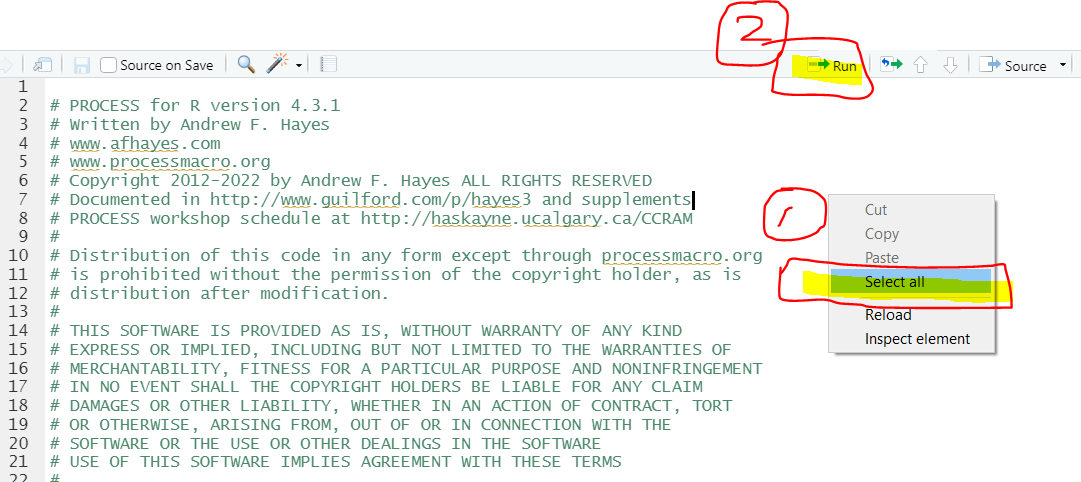
Then, the process() function is in the R environment (You will see the following output). That means that we can use the function.
After setting up the PROCESS in R, we can use Model 4 as a simple example. In the following R code, we first download the data from GitHub. Then, run it using the process().
# Read data from GitHub
data_mediation <- read.csv("https://raw.githubusercontent.com/tidydatayt/mediation_analysis/main/mediation_hypothetical_data.csv")
# run model 4 using PROCESS in R as an example
process(data = data_mediation, y = "Y", x = "X", m ="M", model = 4)
After running the R code above, you should see the following output, which means that you setup PROCESS in R correctly. You can change variable names and the model number to do your own mediation analysis.
********************* PROCESS for R Version 4.3.1 *********************
Written by Andrew F. Hayes, Ph.D. www.afhayes.com
Documentation available in Hayes (2022). www.guilford.com/p/hayes3
***********************************************************************
Model : 4
Y : Y
X : X
M : M
Sample size: 100
Random seed: 611311
***********************************************************************
Outcome Variable: M
Model Summary:
R R-sq MSE F df1
0.2891 0.0836 1.2885 8.9359 1.0000
df2 p
98.0000 0.0035
Model:
coeff se t p
constant 0.6468 0.9247 0.6996 0.4859
X 0.0333 0.0111 2.9893 0.0035
LLCI ULCI
constant -1.1881 2.4818
X 0.0112 0.0553
***********************************************************************
Outcome Variable: Y
Model Summary:
R R-sq MSE F df1
0.9566 0.9150 1.1382 522.3480 2.0000
df2 p
97.0000 0.0000
Model:
coeff se t p
constant 0.0327 0.8712 0.0376 0.9701
X 0.0023 0.0109 0.2093 0.8346
M 2.9318 0.0949 30.8807 0.0000
LLCI ULCI
constant -1.6964 1.7618
X -0.0194 0.0240
M 2.7433 3.1202
***********************************************************************
Bootstrapping progress:
|>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>| 100%
**************** DIRECT AND INDIRECT EFFECTS OF X ON Y ****************
Direct effect of X on Y:
effect se t p LLCI ULCI
0.0023 0.0109 0.2093 0.8346 -0.0194 0.0240
Indirect effect(s) of X on Y:
Effect BootSE BootLLCI BootULCI
M 0.0975 0.0304 0.0368 0.1562
******************** ANALYSIS NOTES AND ERRORS ************************
Level of confidence for all confidence intervals in output: 95
Number of bootstraps for percentile bootstrap confidence intervals: 5000
Discussion